Research News
Study examines how placentas evolved in mammals
By CHARLOTTE HSU
Published July 18, 2022
The fossil record tells us about ancient life through the preserved remains of body parts, like bones, teeth and turtle shells. But how to study the history of soft tissues and organs, which can decay quickly, leaving little evidence behind?
In a new study, scientists use gene expression patterns, called transcriptomics, to investigate the ancient origins of one organ: the placenta, which is vital to pregnancy.
“In some mammals, like humans, the placenta is really invasive, so it invades all the way through the wall of the uterus, into the maternal tissue. In other mammals, the placenta just touches the wall of the uterus. And then there’s everything in between,” says senior author Vincent J. Lynch, associate professor of biological sciences, College of Arts and Sciences.
“So what kind of placentas were early placentas?” he says. “We use gene expression patterns to reconstruct the evolution of the placenta and predict what the placenta of the last common ancestor of eutherian mammals looked like. Our data tells us that this placenta was invasive, and that non-invasive placentas evolved multiple times among mammals. This addresses a 150-year-old mystery. People have been debating what kind of placenta the first one was since then.”
As Lynch explains, all living mammals other than marsupials and egg-laying monotremes are eutherians, which have long pregnancies in which the developing fetus evokes a strong physiological response in the mother.
The research was published on June 30 in eLife. Lynch led the study with first author Katelyn Mika, University of Chicago postdoctoral scholar in human genetics and in organismal biology and anatomy. Camilla M. Whittington and Bronwyn M. McAllan, both at the University of Sydney, are also co-authors.
“Our ability to ask how the placenta might have functioned at different points during its evolution by using the gene expression profiles of currently existing animals to reconstruct the ancestors is a really cool approach and provides us more information on how changing gene expression can contribute to the evolution of a new trait,” Mika says.
To conduct the analysis, the team compared the genes active in the uterus of various mammals during pregnancy. After finding that these gene expression profiles correlated with the degree of placental invasiveness, the scientists used their data to predict what ancestral mammalian placentas looked like.
The study included about 20 species, such as the egg-laying platypus, pouch-bearing marsupials and a range of eutherian mammals that give birth to live young.
The small subset is one limitation of the analysis: The authors write in eLife that research on a larger number of species is needed to help determine the strength of the findings.
Nevertheless, the study makes important contributions in understanding how pregnancy evolved, Lynch says. The results could also benefit modern medicine.
“Knowing which genes are active among different species during pregnancy tells us about how evolution works,” he says. “But it also tells us about what makes a healthy pregnancy, and how things might go wrong. We’re finding the genes that establish the right kind of environment for healthy human pregnancies. If those genes are not expressed in the right way, that might give rise to problems.”
This study was supported by grants from the March of Dimes and the Burroughs Wellcome Fund Preterm Birth Initiative.
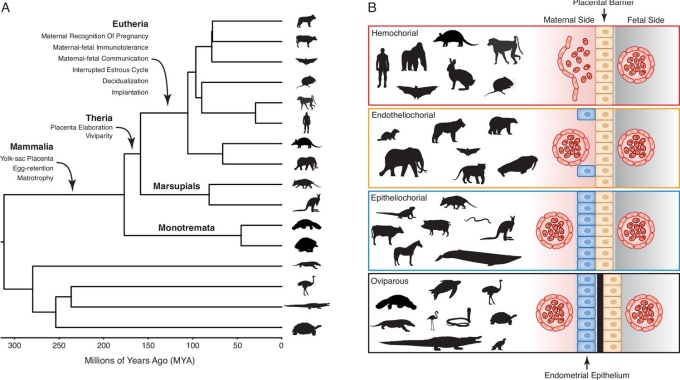
Figure 1. Evolution and arrangement of the maternal-fetal interface and degree of placental invasion in viviparous Amniotes. The paper is here.